GPUMD News
GPUMD-NEP helps publish in "Science", revealing short-range order in semiconductors
Recently, "Science" published the latest research work on short-range order in semiconductor alloys by a collaborative team from Lawrence Berkeley National Laboratory and George Washington University. This work captures and quantitatively characterizes short-range order in semiconductor alloys for the first time in experiments.
Read Article
GPUMD 4.0 paper published in MGE Advances
GPUMD 4.0 was released on April 29, 2025, and the corresponding paper was published online in the domestic journal MGE Advances on August 3, 2025. Bohai University is the first author unit of this paper.
Read Article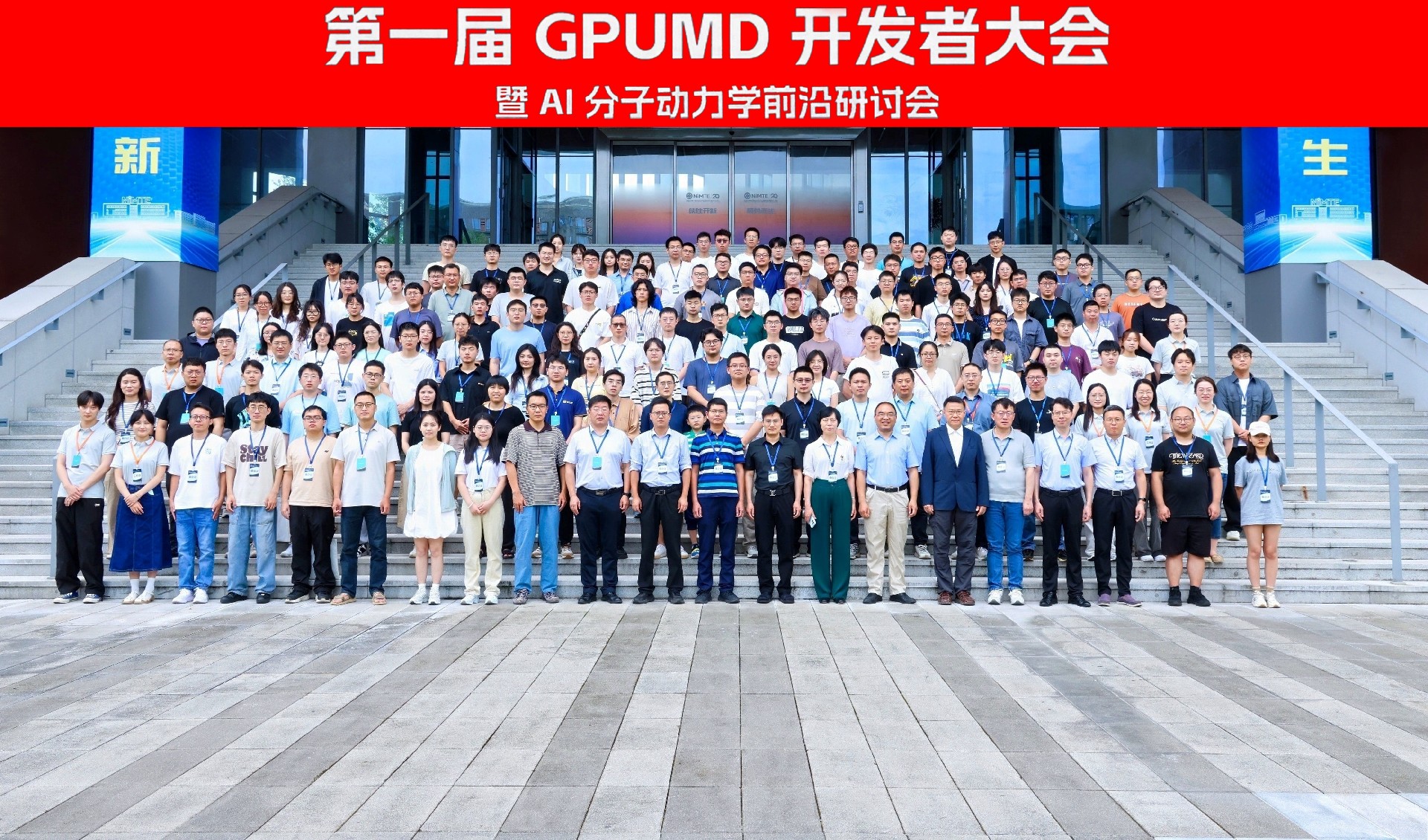
1st GPUMD Developer Conference & AI-MD Frontier Symposium
The 1st GPUMD Developer Conference and AI Molecular Dynamics Frontier Symposium was successfully held in Ningbo in June 2025. The conference gathered developers and experts from all over the country to discuss the latest progress.
Read Article